BrainVoyager v23.0
Advanced Segmentation Tools
The tools described in section Cortex Segmentation and Reconstruction allow to segment the cortex largely automatically by application a pipeline of different anatomical processing tools. From the obtained segmented cortical boundary, surface representations (meshes) can be reconstructed, which are useful for 3D visualizations and cortex-based statistics. Furthermore, topologically corrected meshes can be morphed to create inflated and flattened cortex representations. The folded meshes can also be morphed into spherical representations, which are used for inter-subject cortex-based alignment (CBA). These "standard" segmentation tools can also use the advanced segmentation pipeline described in this section by checking the High-resolution (upsampled to 0.5 mm iso voxel) option in the Automatic Segmentation and Reconstruction dialog; the resulting cortex segmentations will be still have 1mm resolution since the data is upsampled first to 0.5 mm and - after cortex segmentation is complete - downsampled back to 1.0 mm.
While the standard segmentation tools are well-suited to get good cortex segmentations in a couple of minutes, the goal of the advanced segmentation tools described here is to provide highly accurate representations of the inner (white matter / grey matter) boundary (called WM-GM or simply WM subsequently) as well as the outer (pial, grey-matter / cerebro-spinal fluid) boundary (called GM-CSF, pial, or simply GM subsequently) so that they are suitable for measuring cortical thickness in individual brains besides producing more detailed cortex representations. For the implemented cortical thickness measurement method, the WM-GM and GM-CSF boundaries must be determined in volume space. The standard segmentation tools provide only one (nearly) topologically correct segmentation result running close to the WM-GM boundary usually slightly inside grey matter since it is easier to obtain a topologically correct surface when "dilating" the WM-GM border into grey matter. On the other hand, the advanced segmentation tools attempt to estimate the WM-GM and GM-CSF boundaries as precisely as possible so that they can be used as input for cortical thickness measurements. To acheive such high quality segmentations, 1 mm data sets (used for standard segmentation) are not providing a rich enough space to accurately segment the WM-GM and the GM-CSF boundary. One important difference between the advanced and standard segmentation tools is that the advanced tools work on sub-millimeter data sets, preferentially with 0.5-0.8 mm iso-voxel acquisition resolution as used often in studies using ultra-high field (7T and higher) scanners. If only 1mm data sets are available, the first step of the advanced segmentation pipeline upsamples the data to 0.5 mm resolution using sinc interpolation before proceeding, but it is recommended to acquire true (non-interpolated) sub-millimeter data, if possible.
Another important feature of the advanced segmentation tools is that intensity thresholds for segmentation of WM, GM and CSF are calculated adaptively by inspecting local intensity histograms as well as computed gradient fields. Note that the quality of the data submitted to the advanced tools should be very high, otherwise erroneous WM and GM segmentations might result (see Prerequisites in topic Cortex Segmentation). It is also highly recommended to run an automatic intensity inhomogenity preprocessing step before (advanced) cortex segmentation. While the quality of the anatomical 3D data sets should be very high, subsequent measurement of cortical thickness tolerates small topological errors. It is thus usually not necessary to correct the produced result even if there are small errors at focal regions. If one wants to use, however the resulting reconstructions for cortex inflations and cortex-based alignment (CBA), topologically correct segmentations are needed and that may require some manual work. Since BrainVoyager QX 2.8 it is also possible to produce downsampled (1mm) cortex segmentations simplifying manual corrections because of up to 8-fold less voxels (0.5 vs 1mm iso-voixel resolution). Since BrainVoyager 20 it is possible to use the "bridge removal" correction tool also for sub-millimeter data and there are also improved tools for manual segmentations.
The advanced segmentation tools are available in the Advanced Segmentation Tools dialog (see figure below), which can be launched by clicking the Advanced Segmentation Tools menu item in the Volumes menu.
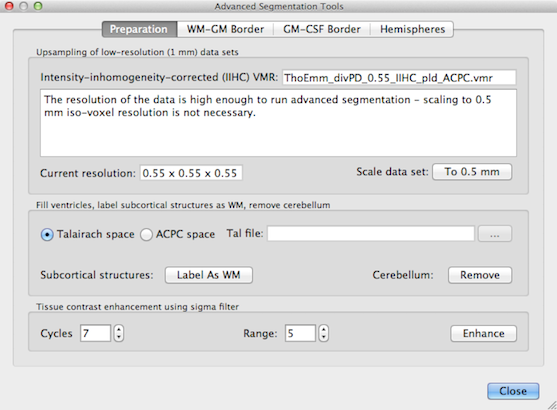
The dialog contains four tabs, Preparation, WM-GM Border and GM-CSF Border and Hemispheres. The Preparation tab contains a set of tools to prepare the original data for advanced segmentation including interpolation to a 0.5 mm resolution, removal of head tissue, labelling of subcortical structures as white matter and tissue enhancement. Some of these tools are built on the standard tools but have been adapted and optimized for a voxel resolution of 0.5 mm. The WM-GM Border and GM-CSF Border tabs contain tools to determine the GM-WM and WM-CSF boundary, respectively.
Since BrainVoyager QX 2.8, the tools work with any (sub-millimeter) voxel resolution (not only in 0.5 mm space as before) by properly adjusting to the resolution and bounding box offset values in the header of VMR data sets.
Copyright © 2023 Rainer Goebel. All rights reserved.