BrainVoyager QX v2.8
The Automatic Cortex Segmentation Pipeline
After loading a 3D data set (VMR) in 1 mm ACPC or Talairach space, you may access the automatic segmentation tools from the 3D Volume Tools dialog. Switch to the Segmentation tab of this dialog and click the Autom. Segm. button. This will invoke the Automatic Cortex Segmentation and Reconstruction dialog (see snapshot below).
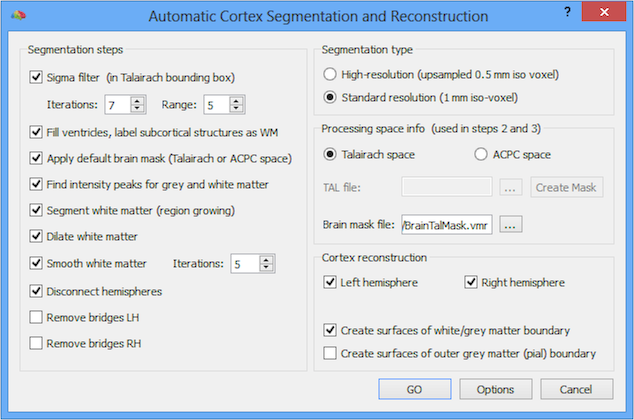
The Segmentation steps field shows the individual steps performed in sequence in voxel space (VMR files), while the Cortex reconstruction field shows the steps performed on surface meshes (SRF files) created from segmented volumes. The Remove bridges LH and Remove bridges RH steps are not selected as default because they require rather long calculations (about 2 minutes per hemisphere on current computers). It is recommended to run the automatic segmentation routine first without these tools in order to check that the obtained results turn out satisfactory. If this is not the case, the tools can be rerun with slightly different settings. If the segmentation turns out satisfactory, you can simply rerun the automatic segmentation tools (eventually with some adjusted parameters) with the bridge removal tool by turning on the respective options.
Since BrainVoyager QX 2.8 it is also possible to run the automatic segmentation pipeline using the advanced segmentation routines by turning on the new High-resolution (upsampled 0.5 mm iso voxel) option in the Segmentaiton type field. When this option is used, the 1 mm ACPC or Talairach data set will be upscaled internally to 0.5 mm and then submitted to various steps of the advanced segmentation tools. When advanced white matter segmentation has been completed, a 1 mm version will be finally computed. From this point onwards, the standard pipeline is used including bridge removal (if enabled) and cortex reconstruction.
The next sections describe the individual steps performed as well as the resulting data files automatically saved to disk.
Copyright © 2014 Rainer Goebel. All rights reserved.