BrainVoyager 20 - A Milestone Release
Flexible subscription-based software licensing
While the hardware licensing (USB dongles) will still be available for some time, the new software-based licensing system is the default licensing scheme for new users and it is also expected that existing users switch to the new system in the coming months. The new licensing system is more convenient to use since it does not require a dongle and thus does not block a free USB port anymore. It also allows more fine-grained subscription-based licenses that better adjusts to the needs and resources of customers. Furthermore, BrainVoyager trial versions are no longer limited to 32-bit Windows but are available now for 64-bit Windows, Mac and Linux operating systems. While license keys are bound to a computer, they can be deleted and moved to another machine. Furthermore, software-based licensing will support floating licenses in the near future.For beginners and experts: The Data Analysis Manager
While BrainVoyager 20 keeps and extends its elegant user interface, a powerful new data analysis management tool has been introduced that supports creation, management and analysis documentation of your projects. The data analysis manager can be used to run basic analysis pipelines (including anatomical and functional preprocessing, alignment, normalization and statistical analyses) with the same parameters for the data of all subjects of a project.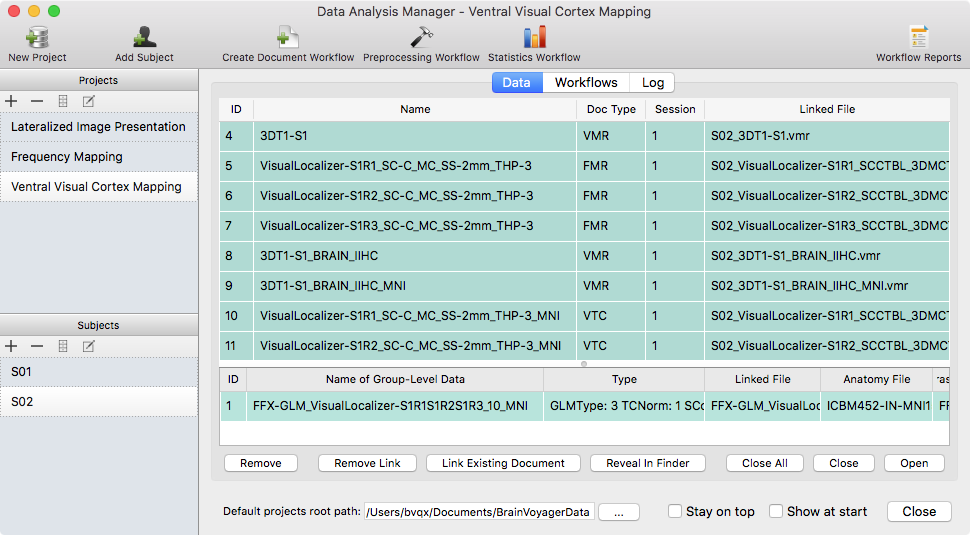
The data manager can thus be viewed as a batch processing tool analyzing big (as well as small) data sets without the need to write scripts. In addition to batch processing, the data manager keeps track of created data and performed analysis steps in a data base providing a persistent overview of the state of analysis of all created experiments. This is not only relevant for beginners but also helps experts to keep track of their projects and to extent analyses with the same workflows as soon as data from new subjects becomes available. You can find a detailed description of the data analysis manager in the User's Guide.
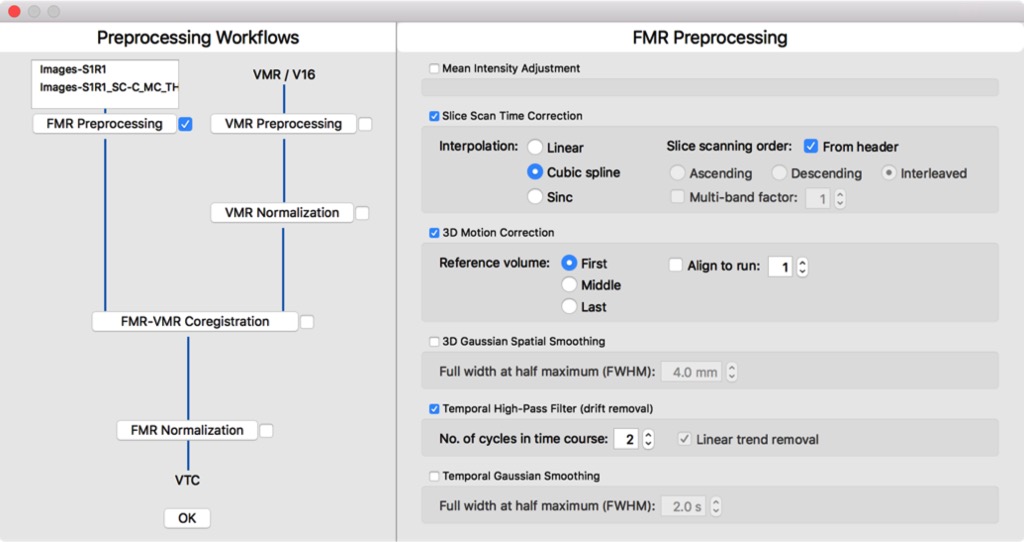
Because of its importance for beginners and experts, I plan to enhance the data analysis manager in several ways in future releases. In the next minor releases, more workflow types will be added, including workflows for multi-voxel pattern analysis, cortex segmentation and cortex-based alignment and statistics. For the next major release, it is planned to allow running workflows not only locally but also on a connected remote compute server or powerful workstation. In this scenario, workflows and input data will be sent to the compute server and results (output data, logs and documentation) will flow back to the local machine for inspection of results.
Automatic MNI Normalization
BrainVoyager 20 introduces automatic MNI normalisation of anatomical and functional documents via standard dialogs for individual data sets as well as for all subjects of an experiment via workflows of the data analysis manager. This was requested by many users for quite some time and it is finally available. The normalization of anatomical (VMR) documents is performed by matching the current VMR to one of the best available template brains resulting in a 12-parameter transformation matrix. While the used template (ICBM-452) is not exactly in MNI space, fixed transformation matrices are appended to the produced matrix to create transformation matrices that transform the input VMR data to the conventional MNI-152 space (default) or to the Colin-27 space. The produced MNI transformation matrix (.trf file) can be used in the "Create VTC" dialog in order to transform functional (FMR) documents to the respective MNI target space. As opposed to a 9 parameter transformation (as used e.g. for ACPC transformation), the MNI transformation matrix uses 12 parameters including also shears in addition to translations, rotations and scales.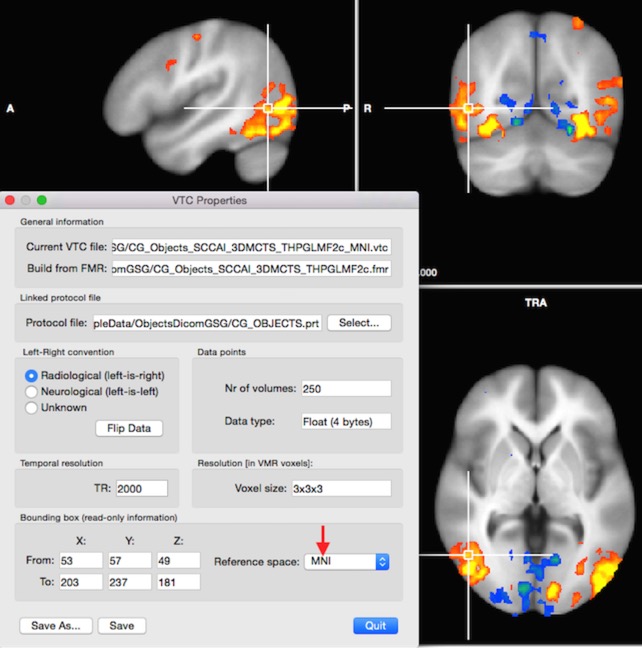
MNI space normalization can be used as an alternative to Talairach space normalization but the choice should be consistent for all data sets belonging to the same experiment, which is ensured when using the data analysis manager. While the two spaces are similar, reported coordinates are slightly different and the used space should be explicitly mentioned when reporting coordinates of activation clusters. In a next update I will present both coordinates (one converted) so that one can get Talairach coordinates when working in MNI space or vice versa.
While the spaces are similar, MNI space normalization has the advantage that the used template-based automatic transformation of VMR data sets operates more robustly and reliably than the automatic Talairach transformation procedure, which makes it better suited for batch processing via scripts or the data analysis manager. It is, however, also possible now to use the template-based approach for Talairach normalization. More details about MNI normalization are provided in the User's Guide.
Representational Similarity Analysis (RSA)
Representational similarity analysis (RSA) is a very useful multivariate tool to analyze the response similarity between evoked distributed fMRI responses in selected regions-of-interest (ROIs). This release provides the possibility to compute representational distance (or dissimilarity) matrices (RDMs) for ROIs including a matrix plot containing color-coded intensity visualizations of the distance measures (e.g. spatial correlation) between pairs of distributed activity patterns from different experimental conditions. Since I find it very helpful to visualize the distance information stored in RDMs in a metric 2D space, I added multi-dimensional scaling (MDS) plots as a default output option when computing RDMs.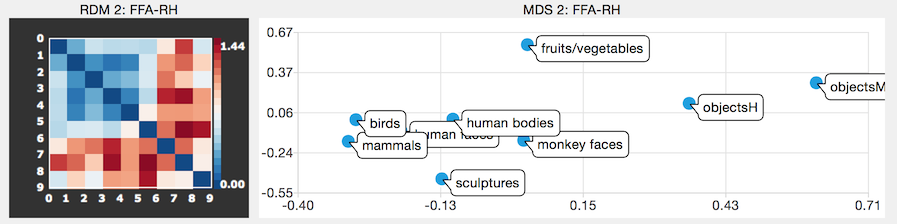
An important aspect of RSA is that it can be applied hierarchically, i.e. the similarity structure between RDMs can itself be assessed by calculating second-level RDMs, i.e. the ordered pairwise first-level dissimilarity values from ROI RDMs are used to calculate a (dis-)similarity measure at the second level. Since second-level comparisons do not require first-level (i.e voxel-level) correspondence, data from other sources can be easily integrated (compared or pooled) at the second-level, including RDMs from multiple subjects, from other measurement modalities and even from computational models. This is an important aspect of RSA since it elegantly solves (or avoids) the necessity to establish first-level correspondence, e.g. between voxels across subjects or between neurons in models and voxels in brains. More details about RSA can be found here.
High-resolution VTC-VTC across-run alignment
BrainVoyager 20 contains also many improvements for analyzing high-resoluion data from ultra-high field scanners with 7 or more Tesla. When scanning sub-millimeter functional data with gradient-echo or spin-echo sequences to measure columnar-like features or to separate responses (roughly) from differet cortical layers, only a small slab of the brain is usually measured as shown in the figure above. For such slab-based sub-milimeter data, the normal across-run FMR motion correction tool in BrainVoyager does not always produce optimal results. Ideally the relevant data from multiple runs within the slab should be aligned very precisely since even small misalignments will strongly reduce the detectability of fine-grained feature or layer-specific activation clusters. In order to perform optimal across-run alignment, a VTC-VTC grid search tool has been introduced in BrainVoyager 20. The term "grid search" indicates that the alignment is not performed by optimization techniques such as gradient descent but simply by exploring transformation parameters in a specified range and by selecting those parameters that produce the best alignment as measured by a goodness-of-fit value that calculates the spatial correlation between the transformed source VTC volume and the target VTC volume. To ensure best results, the region of interest within the slab can be marked to define a mask that restricts the space that will be used when calculating the spatial correlation value. This is a very powerful approach since the alignment will be focused to the most critical region and noisy data (such as infolds and distortions) outside the interesting region will be ignored. Besides VTC-VTC alignment the tool can also be used for local VMR-VTC alignment. Details on how to use this tool can be found in this section of the User's Guide.New Zoom View pane with enhanced manual segmentation tools
While previous versions of BrainVoyager had some manual segmentation tools, I decided to improve these tools for BrainVoyager 20. The tools should be more easily accessible from a centralized place and it should be easier to zoom into small regions than in the past. Say hello to the new Zoom View pane, which allows to show any sub-region of a document that is selected with the mouse.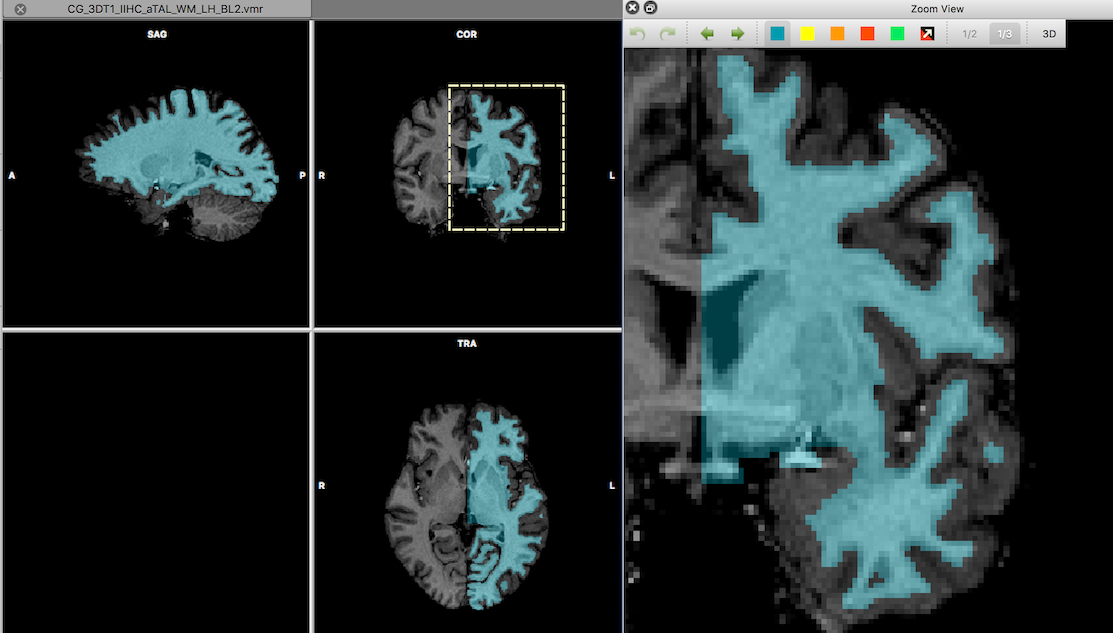
The selected sub-region will be highlighted by a stippled yellow rectangle in the document window and the yellow rectangle can be moved to a new location. For VMR documents, the Zoom View pane shows a toolbar that provides enhanced segmentation tools. Maybe the most important improvement (requested by many users) is the availability of undo/redo operations with an unlimited history of drawing operations. In addition, the toolbar supports pixel-precise drawing, region filling, color selection, toggling between (blended) volumes, and the creation of 3D meshes for all segments (colored labels). For a detailed description of these new tools can be found in the topic Manual Segmentation Tools in the User's Guide.
New Open Docs pane
To switch between documents is easy by clicking on the respective tab bar (in xx mode). With more than 3 open documents, this becomes increasingly difficult since document names are not fully visible anymore. I added the Open Docs pane as a better additional way to switch between open documents by selecting a document by its visual representation in a "cover flow" like layout.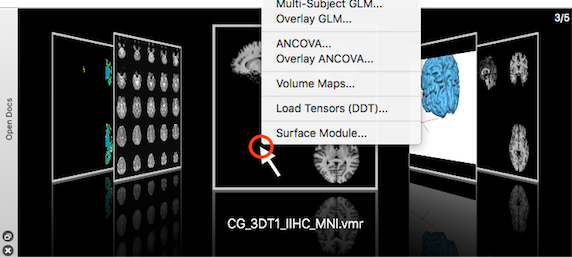
Its also possible to quickly browse through available documents and to select a document by releasing the mouse when the desired document is shown in the central position, together with its full name. To be as useful as document tabs, it is also possible to invoke a context menu from which relevant document-specific functionality can be launched quickly. For more details about the Open Doc pane and other main elements of the user interface, have a look at the Graphical User Interface section in the User's Guide.
A new kid in town: Python
While I am very happy with the C++-based plugin interface that I developed for BrainVoyager, I thought for some time that it should become easier to write plugins that do not require the absolutely highest performance. I considered two languages for integration into BrainVoyager for easier plugin development: Swift and Python. Swift is an amazing new language developed by Apple that quickly replaces Objective C for developing Mac and iOS applications. While the language is elegant and easy to learn, I thought that it will remain a language restricted to software for Apple operating systems and thus would not be usable for cross-platform development. Fortunately Apple has decided to make Swift available as open source in the near future (as announced in June 2015) and I may consider the language for cross-platform development at a later time. In my view the best established cross-platform programming language for "easy" scientific software development is Python. As an interpreted language, Python allows scripting and fast prototyping in an interactive shell or console as well as running object-oriented program files. Extension modules such as NumPy and SciPy make Python a high performance language for scientific computing. Combined with the interactive interpreter these features have made the language very popular for scientific programming and prototyping rivalling Matlab in many ways.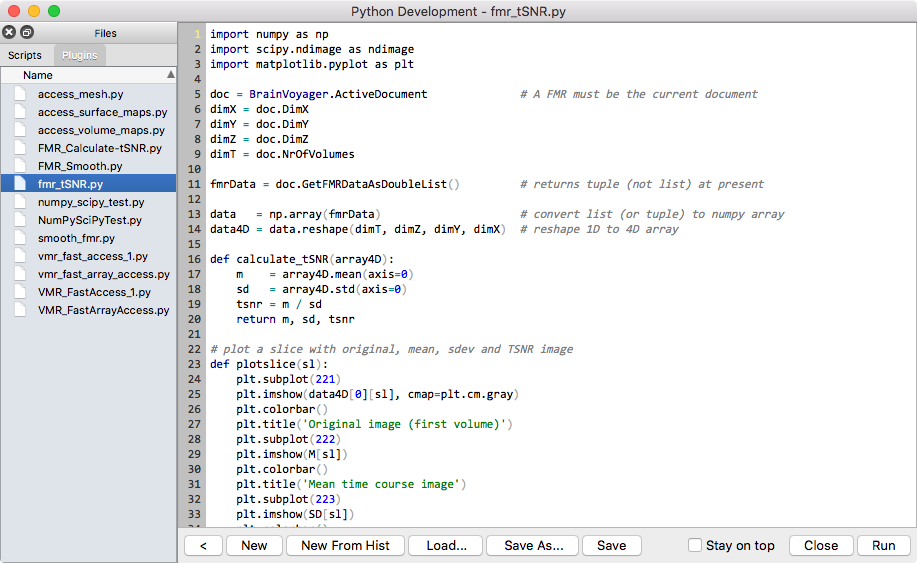
Another important property of Python is that the interpreter can be embedded in C++ programs, which was ideal for my goal to add a language for interactive exploration, scripting and plugin development. The PythonQt project even provides a direct interface to the cross-platform Qt libraries that I use for cross-platform GUI development right from the embedded Python interpreter; this establishes a very easy and powerful way to build GUI-driven scripts and plugins using the same language as opposed to a mix of C++ and JavaScript that is needed for the standard plugin development framework. Besides GUI development using Qt, it was also easy with PythonQt to inject a "BrainVoyager" object into the Python interpreter so that Python code can call the same commands as the standard JavaScript scripting interface, i.e. it offers a complete BrainVoyager scripting alternative. To see how easy it is to script BrainVoyager from Python, have a look at the provided sample scripts located in the "BVExtensions/PythonScripts" folder. In order to also serve as a language for plugin development, more hard work was needed that required diving into Python extension modules. To provide numpy array access, I implemented a C extension module called "bv" that allows to access internal BrainVoyager C pointers that reference important data structures in memory (e.g. VMR volumes, VTC volumes, mesh arrays, volume and surface maps). Since numpy arrays support vectorized operations avoiding explicit loops (like matrices in Matlab), Python code using numpy arrays can reach very high performance levels that come close to C/C++ code despite being executed by the Python interpreter. Have a look at the provided examples in the "BVExtensions/PythonPlugins" folder to learn from simple examples how easy it is to create (GUI) plugins in Python.
While the current implementation already provides a solid basis for using Python within BrainVoyager, Python support is marked as experimental in BrainVoyager 20.0 since some features are missing for a complete integrated development environment, most notably debugging support and a more powerful multi-document editor. Furthermore, the latest Python 3.x version is not supported but only Python 2.7. This is partially due to the fact that Mac OS X comes with Python 2.7 installed and BrainVoyager uses the system implementation resulting in robust Python support on Apple computers without the need to manually install Python. To be consistent across platforms, version 2.7 of Python 2.7 must also be installed on Windows, which does not come with a preinstalled Python interpreter. For users that are not interested in using Python, a version of BrainVoyager 20.0 without Python is available that does not require installation of Python. The Linux version of BrainVoyager 20.0 does not come with Python support but it will be added in an upcoming minor update of BrainVoyager 20. Since Python is so useful for data exploration, scripting and plugin development, the created BrainVoyager Python environment will improve quickly in subsequent program updates.
I hope you will enjoy BrainVoyager 20 providing analysis and visualization tools that are useful for beginners and experts alike. The next releases will further improve on the foundation laid in this release. As always, our goal is to provide the most elegant, easy-to-use and comprehensive software package to perform advanced brain imaging data analyses and to produce beautiful images and animations of achieved results. If you want to get a trial version or upgrade an existing license, please consult this page for details.