BrainVoyager QX 2.0 - Episode 5: Recursive Feature Elimination
June 13, 2009
In the last blog entry, we have seen how SVMs can be applied to fMRI data extracted from regions-of-interest (ROIs). Using ROIs to limit the voxels included in training a classifier has both a conceptual and a technical benefit. Conceptually, it allows to ask the specific question whether a particular brain region (e.g. the fusiform face area) has the ability to discriminate multivariate patterns of interest (e.g. patterns evoked from several different views of two faces). As a technical benefit, the ROI approach reduces the number of features (voxels) and, hence, helps to avoid the “curse of dimensionality”.
Multivariate Brain Mapping
As discussed in episode 2, the ROI approach is, however, not always suitable since one might not know the locations a priori where different conditions can be separated using MVPA tools. In these situations, one would like to find those brain regions, which are able to discriminate two conditions, i.e. one is interested in multivariate brain mapping. One interesting multivariate brain mapping strategy performs a multivariate analysis at each voxel including the responses from voxels in the local neighborhood. This “searchlight” approach is described in more detail in a later blog entry. In this blog the focus is on recursive feature elimination (RFE), which is another multivariate brain mapping approach. RFE allows to detect (sparse) discriminative patterns that are not limited to the local neighborhood but that may be spread across the whole brain. The basic principle of RFE is to include initially all voxels of a large region, and to exclude voxels, that do not contribute in discriminating patterns of different classes. Whether a voxel in the current feature set contributes enough to be kept is determined by the weight value of a voxel resulting from training a classifier (e.g. SVM) with the current set of features. in order to increase the likelihood that the right voxels are selected, feature elimination progresses gradually and includes cross-validation steps. In each feature elimination step, a small proportion of voxels is discarded until a core set of voxels remains with the highest discriminative power. Note that using SVM to separate “good” from “bad” voxels implements a multivariate feature selection strategy as opposed to univariate feature selection using single-voxel F or t values from a statistical analysis. Nonetheless, an initial feature reduction step using a univariate method might be useful if one wants to restrict RFE to the subset of “active” voxels.RFE Details
The RFE implemented in BrainVoyager QX 2.0 is based on DeMartino et al. (2008, NeuroImage, 43, 44-48). It includes two nested levels of cross-validation to maximize the chance to keep the “best” voxels. At the first level, the training data is partitioned in NF folds and RFE is applied NF times. In each application, one of the folds is put aside for testing generalization performance while the other folds together form the training data for the RFE procedure, i.e. for each of the NF RFE’s another “split” of the data is used. When all separate RFE’s have been performed, the final generalization performance is determined as the average of the performance across the NF different splits, separately for each reduction level (see below). The final set of voxels (for a specific reduction level) are obtained by merging the voxels with the best weights (highest absolute values) across all splits.The training data from each first-level split is used for a separate RFE procedure while the fold with the test data is set aside and only used for performance testing. The training data is then partitioned again in L sub-folds and a SVM is trained on L splits in order to obtain robust weight rankings for feature elimination. A voxel’s ranking score is obtained by averaging the weights of that voxel across the different second-level splits. The absolute values of these scores are then ranked and the voxels with the lowest ranks are removed. The “surviving” voxels are then used for the next RFE iteration, which starts again with (a new) partitioning of the data in L folds. The whole procedure is repeated R times until a desired number of voxels has been reached. As described above, the RFE level producing the highest generalization performance across all first-level splits is finally selected and the level’s set of voxels is determined by merging the best voxels of the respective first-level splits.
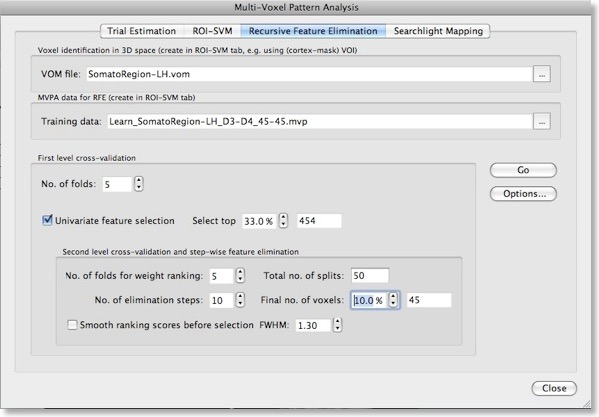
Application
The figure above shows the “Recursive Feature Elimination” tab of the “MVPA” dialog in BrainVoyager QX 2.0. To perform RFE, we need to provide as input a VOM and a MVP file. As described in a previous blog entry, a VOM is similar to a VOI with the exception that voxels are stored in the resolution of functional data (VTCs/VMPs) instead of the resolution of the “hosting” anatomy (VMR); furthermore a VOM may contain for each voxel a floating-point value (e.g. a statistical map value or a SVM weight). The VOM file can be created from a VOI in the ROI-SVM tab or using the new “Create VOM” dialog (“Options > Create VOI Map” menu item). Starting from a VOM (derived from a VOI) allows to determine the size of the brain region to be used for recursive feature elimination. While RFE can be applied to the whole brain, it is often more appropriate to select a large interesting region, such as the visual cortex or the frontal lobe. When such restrictions are not desired, it is best to convert a cortex mask VOI to a VOM. Use the “Browse” button on the right side of the “VOM file” text field to specify the desired brain region.The second input needed for RFE is a MVP file containing the training data for the initial (full) set of voxels corresponding to those in the selected VOM file. The easiest way to get both the VOM and the desired MVP file is to use the “ROI-SVM” tab, which allows to extract estimated BOLD responses for any region of interest as described in episode 3. If trial estimates for the desired data is not yet available, you must first estimate them from a set of selected VTC files (“Trial Estimation” tab) resulting in a set of VMP files containing the trial estimates per voxel (see episode 2). When clicking the “Create” button in the “” field of the “ROI-SVM” tab, a specified VOI is first converted in a VOM to get the voxels in native resolution. The VOM voxels are then used to extract the trial responses from the VMP files (for two specified classes). The result is stored in a MVP file in form of a “trial responses x voxels” matrix (see plots below). Note that a VOM file forms an important link between the voxels in the MVP matrix and the location of the voxels in the brain: the i’th entry in a MVP file contains trial estimates but no voxel coordinates while the i’th entry in a VOM file contains the voxel’s x, y and z coordinates. Because of this important link, the program creates a MVP file and a VOM file with the same name when extracting trial estimates from VMP files. In the used “somatosensory” example data, the VOM name (and original VOI name) is “SomatoRegion-LH” and this name appears also in the selected MVP file (see above) indicating that both files refer to the same voxels.
With the input specified, the RFE procedure can be started by clicking the “GO” button. Before that, you may want to review and eventually change some options. The number of first-level folds (NF) can be specified using the “No. of folds” spin box in the “First level cross-validation” field, which is set to “5” as default. While RFE implements a multivariate feature selection strategy, it may be useful to restrict the number of voxels to some extent using a univariate strategy. If you would like to add univariate selection prior to RFE (performed for each first-level split), turn on the “Univariate feature selection” option and specify the desired percentage of surviving voxels using the “Select top percent” spin box. When you change the percent value, the number of remaining voxels are previewed in the text field on the right side of the percent spin box.
Since RFE uses itself a cross-validation scheme, the “Second-level cross-validation and stepwise feature elimination” field is visually nested within the “First level cross-validation” field. At each RFE level (see below), the data is partitioned again in a number of folds (Nx), which can be adjusted using the “No. of folds for weight ranking” spin box. For each of the created splits, a supprt vector machine is trained and the weights are ranked according to their absolute value. The voxels with the highest weights across all splits are selected. The voxels are, however, not reduced to the final number in one step, but proceeds stepwise resulting in several “RFE levels”. The number of RFE levels can be specified by changing the “No. of elimination steps” spin box (default: 10). Using such a stepwise procedure should help, like the cross-validation approach, to increase the robustness in finding the “best” voxels. The percentage of target voxels, which should remain after running through all RFE levels can be specified in the “Final no. of voxels” spin box. The text field on the right side of this spin box shows the corresponding number of voxels. Note that If the univariate selection option is enabled, the specified percentage is relative to the number of voxels filtered by univariate feature elimination.
As a final option, it is possible to slightly smooth spatially the obtained ranking values prior to selection. This step will introduce a bias towards favoring spatially clustered regions over isolated voxels. In some applications, this smoothing step proved beneficial.
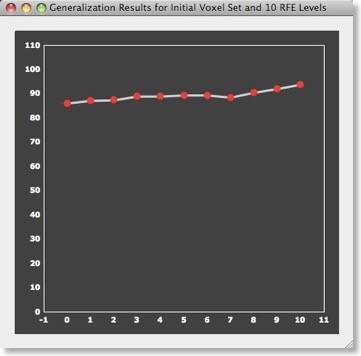
After running RFE, you will see a plot showing the average generalization performance assessed with the separated test data sets for each RFE level. While not always the case, the generalization performance often increases while decreasing the number of voxels. A slight increase is, for example, observed for the sample data set (see plot above) when comparing D2/D4, D2/D3 or D3/D4. Note that because of random generation of folds, you will not get exactly the same results (nor exactly the same voxels) if you re-run the procedure with the same settings.
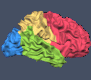
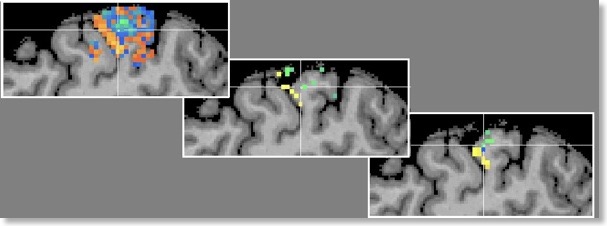
Besides the generalization performance plot, the RFE will also show the resulting voxels and associated weights by visualizing the resulting VOM as a volume map (VMP). The figure on top of this blog shows the SVM weights obtained when comparing conditions D2 and D4 for a sagittal slice with the original (“full”) voxel set and after RFE with a final number of 68 voxels (no univariate selection). The middle plot shows the result when smoothing was turned off, while the plot on the right side shows the result when smoothing was used with a FWHM value of 1.4. While already visible with the full voxel set, RFE seem to have indeed found voxels separating the two digits D2 and D4. While discrimination of D2/D3 and D3/D4 worked almost equally well, the resulting voxels were clustered, which is expected since the representations of neighboring digits will be much more overlapping.
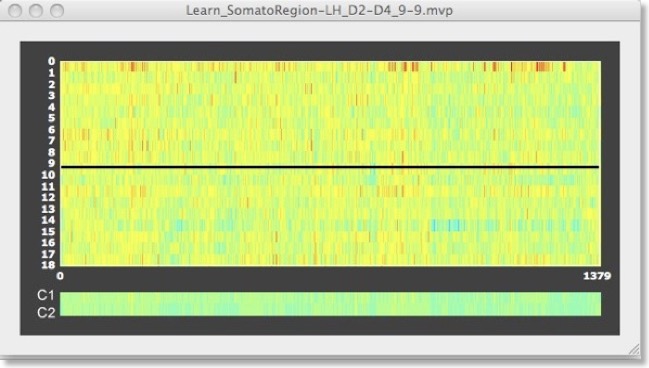
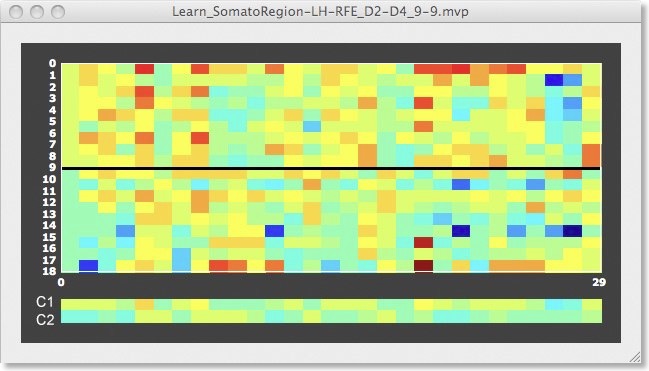
With the help of the voxel extraction feature from VOMs in the “ROI-SVM”, it is also possible to show the trial response data for each voxel of the final RFE level. The top panel in the figure above shows the D2/D4 data for one run (9 trials per class) for the original set of voxels (1378). The black line separates the 9 trials of the D2 and D4 class. The two “lines” at the bottom of the plot show an average of the voxels’ responses separately for each class, which look very similar. The panel below shows as selection of that data for the voxels identified by recursive feature elimination (from 1378 to 28 voxels). This plot reveals that the voxels selected are indeed “good” voxels since many voxels have a positive value for class 1 (orange-red color) but a negative value for class 2 (blue-green color).